12.3. hplc-py#
import numpy as np
import matplotlib.pyplot as plt
from molass_data import SAMPLE1
from molass.DataObjects import SecSaxsData as SSD
ssd = SSD(SAMPLE1)
x, y = ssd.xr.get_icurve().get_xy()
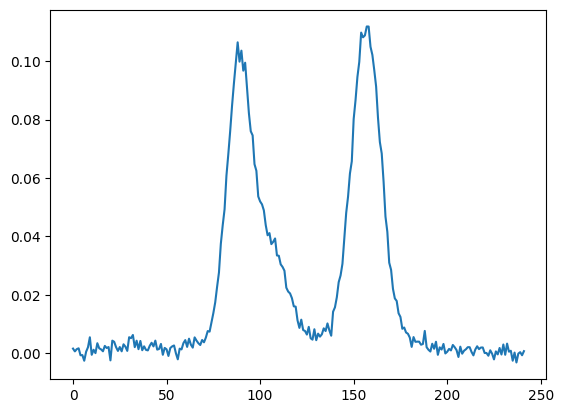
fig, ax = plt.subplots()
ax.plot(x, y)
plt.show()
csv_path = "hplc-py-example.csv"
np.savetxt(csv_path, np.column_stack((x/20, y)), delimiter=",", header="time,signal", comments='')
from hplc.io import load_chromatogram
from hplc.quant import Chromatogram
example = load_chromatogram(csv_path, cols=['time', 'signal'])
chrom = Chromatogram(example)
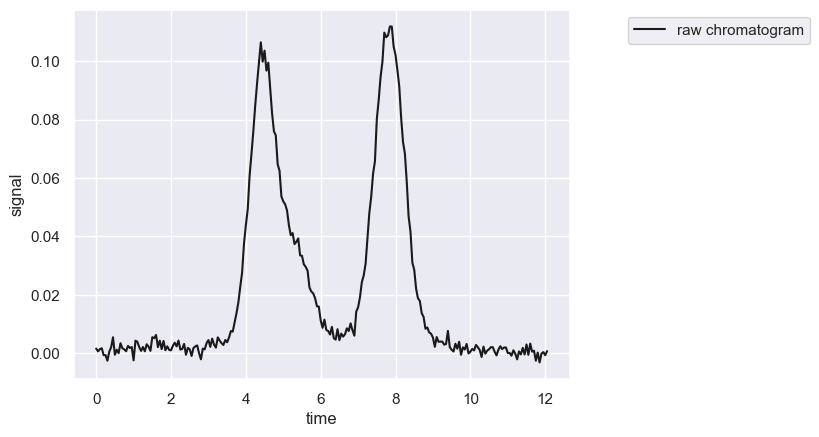
chrom.show()
[<Figure size 640x480 with 1 Axes>, <Axes: xlabel='time', ylabel='signal'>]
peaks = chrom.fit_peaks(prominence=0.05)
peaks.head()
Performing baseline correction: 100%|██████████| 49/49 [00:00<00:00, 5891.21it/s]
Deconvolving mixture: 100%|██████████| 1/1 [00:02<00:00, 2.08s/it]
| retention_time | scale | skew | amplitude | area | signal_maximum | peak_id | |
|---|---|---|---|---|---|---|---|
| 0 | 0.4 | 0.011160 | 0.143289 | 0.001278 | 0.007779 | 0.005471 | 1 |
| 0 | 0.7 | 0.012835 | 0.413577 | 0.000458 | 0.004088 | 0.003025 | 2 |
| 0 | 0.8 | 2.944615 | 9.217561 | 0.012458 | 0.249099 | 0.003237 | 3 |
| 0 | 4.6 | 0.407060 | -2.034294 | 0.067984 | 1.359681 | 0.099460 | 4 |
| 0 | 4.6 | 0.784367 | 5.773530 | 0.054106 | 1.082119 | 0.050649 | 5 |
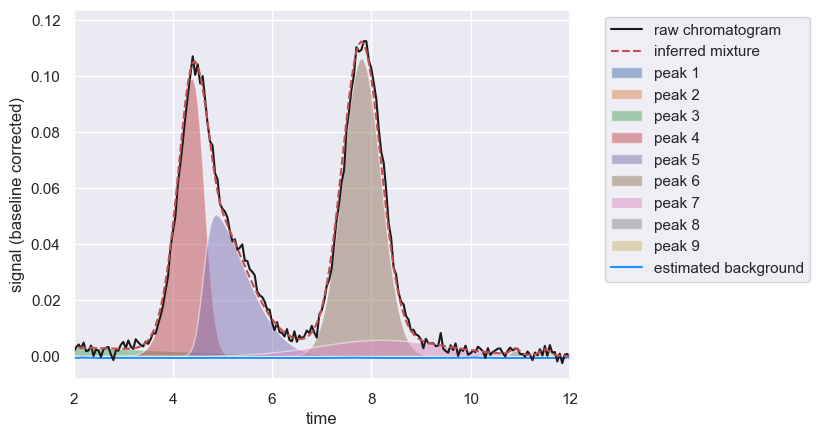
chrom.show([2, 12])
[<Figure size 640x480 with 1 Axes>,
<Axes: xlabel='time', ylabel='signal (baseline corrected)'>]